Base composition and proximity of dnaA and dnaN genes were used by Fraser et al. (Science 281:375-388, 1998) to establish an origin of replication and the coordinate system used herein. TP0001 is the predicted dnaA gene and TP0002 is the predicted dnaN gene; TP0001 starts at coordinate 4. Between TP0001 and the last ORF in the circular genome, TP1041, a predicted ClpP gene, a region of transcriptional divergence is hypothesized; genes transcribed in the right direction (direction of replication) may ha ve a higher translational efficiency (Fraser et la., Ibid).
The average G + C content of the genome is 52.8%. The codon usage as presented on the CUTG Web site (http://www.kazusa.or.jp/codon/) (Nakamura, Y., Gojobori, T. and Ikemura, T. (1997) Nucl. Acids Res. 25, 244-245.) is as follows (based on 29578 codons):
fields: [triplet] [frequency: per thousand] ([number])
UUU 29.3( 867) UCU 18.2( 538) UAU 13.7( 405) UGU 7.3( 216) UUC 13.1( 388) UCC 9.6( 283) UAC 13.6( 402) UGC 6.1( 181) UUA 8.1( 241) UCA 7.4( 220) UAA 0.8( 24) UGA 0.8( 23) UUG 21.6( 638) UCG 10.9( 322) UAG 1.2( 36) UGG 7.8( 231) CUU 22.6( 669) CCU 11.5( 341) CAU 9.0( 267) CGU 19.7( 584) CUC 16.8( 498) CCC 6.9( 205) CAC 10.8( 320) CGC 16.5( 489) CUA 4.9( 146) CCA 4.8( 141) CAA 11.6( 342) CGA 5.1( 152) CUG 19.9( 590) CCG 10.7( 317) CAG 28.6( 846) CGG 10.9( 322) AUU 26.0( 770) ACU 11.7( 347) AAU 16.5( 488) AGU 11.6( 344) AUC 18.9( 560) ACC 12.4( 366) AAC 12.9( 381) AGC 9.9( 293) AUA 8.6( 255) ACA 9.1( 270) AAA 16.3( 482) AGA 5.2( 155) AUG 24.6( 728) ACG 17.5( 517) AAG 30.2( 892) AGG 7.2( 213) GUU 23.4( 693) GCU 17.0( 503) GAU 31.9( 944) GGU 22.7( 672) GUC 12.8( 378) GCC 13.3( 392) GAC 20.7( 613) GGC 13.4( 397) GUA 15.5( 457) GCA 27.9( 824) GAA 27.3( 808) GGA 16.7( 493) GUG 39.7( 1173) GCG 40.5( 1198) GAG 36.2( 1070) GGG 22.2( 658)
Current analyses of this database show 555 identifiable proteins and 177 conserved but uncharacterized proteins; the remaining ORFs are hypothetical. The determination of ORFs is outlined in Fraser et al., Science 281:375-388, 1998. Taking the sequence s from GenBank, each predicted protein has been analyzed herein using gapped BLAST and Psi-BLAST to capture overall similarities and BLOCKS and ProDom to discern shared domains (Documentation). Of the 555 identified proteins, 242 have been assigned an EC number. A few ORFs with inherent frameshifts have been encountered; in these cases, we provide the original "uncorrected" sequence as well as a "corrected" sequence.
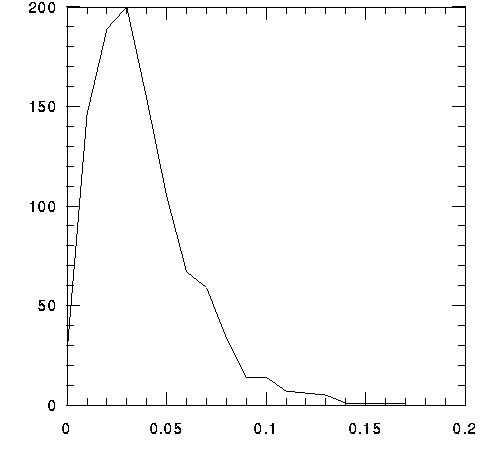
The average molecular weight (length) of the 1041 proteins is 37.6 kDa (340 aa); the median molecular weight (length) is 32.3 kDa (292 aa). The average and median estimated pI values are 8.36 and 8.32 respectively. These estimates differ slightly from estimates reported by Fraser et al. (Ibid, 1998).

x 1000 KDa
Of the 1041 predicted proteins, 160 appear to be secreted, 259 have transmembrane domains, 119 have coiled-coil regions as determined by suite of structural analysis programs in the SEALS package available from NCBI/NIH. Of the 160 proteins predicted t o have a signal peptide, 27 had a best hit (by criteria explained in the following paragraph) to E.coli proteins, whereas 28 had a best hit to B.subtilis proteins. The majority of secreted proteins are not that similar to either E.coli or B.subtilis. Henc e signal peptide prediction for T.pallidum was based on both gram positive and gram negative models. Of the 1041 proteins, 170 have similarity to a PDB database sequence using BLAST2. A hit requires an E value of 0.0001 or better (lower). A higher fractio n could undoubtedly be shown to be homologous to PDB sequences using Psi-BLAST and filtering (Huynen, M. et al., J.Mol.Biol., 280:323-326, 1998).
When T.pallidum proteins are compared to E.coli and B.subtilis proteins, using BLAST2, 174 are found to have a better hit to E.coli proteins, whereas 308 have a better hit to B.subtilis proteins, consistent with the finding that Treponemas are neither gram-positive nor gram negative. In this analysis, the E value for the best hit to a genome must be 100 times smaller (better) than the E value for the second genome. Also, only E values of 1e-03 and lower are considered. Note that approximately 250 of th e identified and conserved T.pallidum proteins (732) do not have a best hit to one of these bacteria by these criteria. Best hits for E.coli and B.subtilis fall to 135 and 238 when archaeal and eukarotic sequences are included in the study, representing 72 and 53 hits respectively.
When T.pallidum proteins are compared to C.trachomatis and M.genitalium proteins, 71 have a best hit to M.genitalium and 304 have a best hit to C.trachomatis, using the criteria stated above (E value of 1e-03 or better and E value 100 times or more sma ller than the next best hit). This trend of greater similarity to C.trachomatis holds for secreted proteins, with 23 best hits to C.trachomatis and merely 4 to M.genitalium (out of 160 T.pallidum secreted proteins).
The original concept of paralog contained the notion that functional differences would evolve after duplication -- that paralogs were not orthologs. In the absence of biochemical data, sequence analysis can't be certain that paralogous relationships wi ll be based on different functions, hence the word paralog is used in a loose sense herein and in the literature to denote similar proteins that are not thought to be orthologous. When duplicated genes with the same activity are being discussed, we will t ry to use the term isolog, leaving the term ortholog for proteins in different genomes. When paralog is used in a strict sense, different functional activities are implied. 299 of the 1041 predicted proteins in T.pallidum have similarity to one or more pr oteins in the organism, using a BLAST2 cutoff E value of 1e-03. Similarities are reported in the Paralog Field irrespective of whether they may be true paralogs or isologs.