Herpesvirus |
|---|
| Introduction |
|---|
| Herpesviridae is the name of a family of enveloped, double-stranded DNA
viruses with relatively large complex genomes. They replicate in the
nucleus of a wide range of vertebrate hosts, including eight varieties
isolated in humans, several each in horses, cattle, mice, pigs, chickens,
turtles, lizards, fish, and even in some invertebrates, such as oysters.
Human herpesvirus infections are endemic and sexual contact is a significant
method of transmission for several including both herpes simplex virus 1 and 2 (HSV-1,
HSV-2), also human cytomegalovirus (HHV-5) and likely Karposi's sarcoma herpesvirus
(HHV-8). The increasing prevalence of genetial herpes and
corresponding rise of neonatal infection and the implication of Epstein-Barr virus
(HHV-4) and Karposi's sarcoma herpesvirus as cofactors in human cancers
create an urgency for a better understanding of this complex, and highly
successful virus family. |
| Virion Structure |
|---|
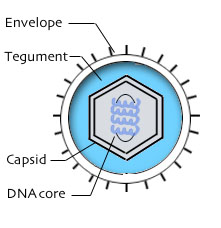
All herpesvirus virions have four structural elements.
|
| Genome Characteristics |
|---|
| Herpesvirus genomes range in length from 120 to 230 kbp with base
composition from 31% to 75% G+C content and contain 60 to 120 genes.
Because replication takes place inside the nucleus, herpesviruses can use
both the host's transcription machinery and DNA repair enzymes to support a
large genome with complex arrays of genes. Herpesvirus genes, like the
genes of their eukaryotic hosts, are not arranged in operons and in most
cases have individual promoters. However, unlike eukaryotic genes, very few
herpesvirus genes are spliced. The genes are characterized as either essential or dispensable for growth in cell culture. Essential genes regulate transcription and are needed to construct the virion. Dispensable genes for the most part function to enhance the cellular environment for virus production, to defend the virus from the host immune system and to promote cell to cell spread. The large numbers of dispensable genes are in reality required for a productive in vivo infection. It is only in the restricted environment of laboratory cell cultures that they are dispensable. All herpesvirus genomes contain lengthy terminal repeats both direct and inverted. There are six terminal repeat arrangements and understanding how these repeats function in viral success is an interesting part of current research. |
| Biological Properties |
|---|
Four biological properties characterize members of the Herpesviridae family.
|
| Strategies for Success |
|---|
| The success of herpesvirus infections depends upon several strategies. The
first is the fast efficient way the virion invades the host cell, turning
off host protein synthesis and releasing viral DNA into the nucleus, where
replication and virion production start immediately. Another strategy that
herpesviruses share is the ability to thwart attacks from the host. Tactics
include inhibiting splicing of mRNA, blocking presentation of antigenic
peptides on the cell surface and blocking the apoptosis (cell death) induced
by viral gene expression. A third important strategy shared by
herpesviruses is their ability to hide their bare, circularized genome in
the nucleus of lymphoma and central nervous system cells and then return to
productive infection months, even years later. These latent herpesvirus
infections are often benign, but can be devastating to newborns and
immuno-suppressed individuals. |
| Herpesviridae Subfamilies |
|---|
| Alphaherpesvirinae. Members of this subfamily are neurotropic (infect nervous system tissue), have a short reproductive cycle (~18 hr.) with efficient cell destruction and variable host range. The human Alphaherpesvirinae with their commom name, scientific name and the disease they cause are: |
| Herpes simplex virus 1 | Human herpesvirus 1 | facial, labial and ocular lesions |
| Herpes simplex virus 2 | Human herpesvirus 2 | genital lesions |
| Varicella-zoster virus | Human herpesvirus 3 | chickenpox and shingles |
| Betaherpesvirinae. Members are lymphotropic, have a long reproductive cycle, restricted host range and infected cells become enlarged (cytomegalo). Human Betaherpesvirinae include: |
| Human cytomegalovirus | Human herpesvirus 5 | infectious mononucleosis |
| (no common names) | Human herpesvirus 6 & 7 | mild early childhood roseola |
| Gammaherpesvirinae. These herpesviruses are also lymphotropic and specific for either T or B lymphocytes. Members of this subfamily isolated in humans are: |
| Epstein-Barr virus | Human herpesvirus 4 | cofactor in human cancers |
| Karposi's sarcoma herpesvirus | Human herpesvirus 8 | cofactor in Karposi's sarcoma which was extremely rare until the advent of AIDS. |
| Research, Vectors & Bioengineering |
|---|
| Recent research has uncovered the function or multiple functions of many herpesvirus genes, while some elude understanding and remain the focus of intense study. Other areas of current and future research include understanding the mechanisms for establishing, maintaining and reactivating latency, interactions between viral and host proteins and details of the variety of mechanisms involved in gene regulation. |
| Herpesviruses are potential vectors for application to several problems in human health. HSV-1 is a good candidate because it is possible to replace large segments of the genome with genes of choice. It's affinity for sensory nerves allows it to target gene therapy to the central nervous system, while it's capacity for cell destruction could be a powerful weapon for selective destruction of cancer cells. An HSV-1 vector could be engineered to express antigens that induce immunity against a variety of infectious agents. Designing such vectors will require profound understanding of the herpesvirus genomes. |
| General References |
|---|
| Three recent publications contain both general and comprehensive reviws of the Herpesviridae family: |
|
Sexually Transmitted Diseases, third edition (1999), ed. Holmes et al., pub. McGraw Hill. Peter E. Pertel, Patricia G. Spear, chapt. 20, "Biology of Herpesviruses", Lawrence Corey, Anna wald, chapt. 21, Genital Herpes", W. Lawrence Drew, Michael P. Bates, chapt. 22, "Cytomegalovirus Infection", and Karen S. Slobod, John W. Sixbey, chapt 23, "Epstein-Barr Virus Infection" |
| Herpesvirus: Genetic variability and recombination, Kenichi Umene, (1998), pub. Touka Shobo. |
|
Fields Virology, third edition (1996), ed. Fields, Knipe, Howley, pub. Lippincott, Williams & Wilkins. Bernard Roizman, chapt. 71, "The Family Herpesviridae: A brief introduction", and Bernard Roizman, Amy E. Spears, chapt. 72, "Herpes Simplex Viruses and Their Replication" |
| Two interesting web sites: |
|
Edward K. Wagner at University of California, Irvine, http://darwin.bio.uci.edu/~faculty/wagner/ University of Rochester Medical Center, Intro. to Virology, Herpesvirus (HSV-1, EBV, KSHV) http://www.urmc.rochester.edu/smd/mbi/grad2/pdf/ |
|
Bernard Roizman The Marjorie B. Kovler Oncology Laboratries University of Chicago 910 East 58th Street Chicago, IL 60637 Nina Thayer Biosciences Division Los Alamos National laboratory Los Alamos, NM 87544 |

Curator: Bernard Roizman Annotator: Nina Thayer |